![PDF] Rank–rank hypergeometric overlap: identification of statistically significant overlap between gene-expression signatures | Semantic Scholar PDF] Rank–rank hypergeometric overlap: identification of statistically significant overlap between gene-expression signatures | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/85b11130c3d0fcb217dc7f1976ed6c909ac6d4b8/3-Figure1-1.png)
PDF] Rank–rank hypergeometric overlap: identification of statistically significant overlap between gene-expression signatures | Semantic Scholar

Immunohistochemical and quantitative RT-PCR methods to assess RANK expression in normal and neoplastic canine mammary gland - Raquel Sánchez-Céspedes, Yolanda Millan, Silvia Guil-Luna, Jesús García-Macías, Lorella Maniscalco, Selina Iussich, Raffaella ...

Framework of TSNBA. PPI network and gene expression data are integrated in the interaction activity matrix to rank genes for their relevancy to the perturbation.
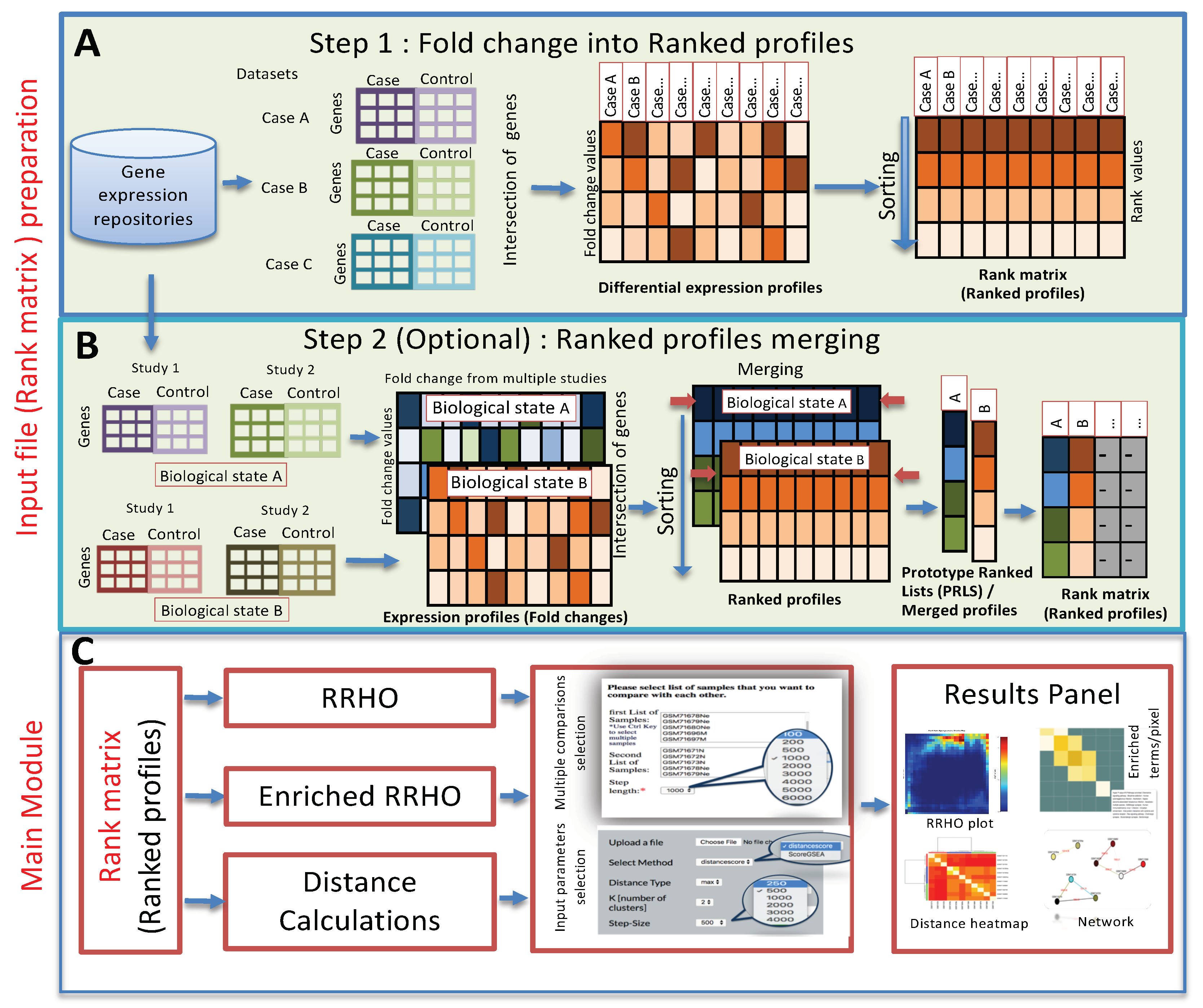
IJMS | Free Full-Text | RankerGUI: A Computational Framework to Compare Differential Gene Expression Profiles Using Rank Based Statistics
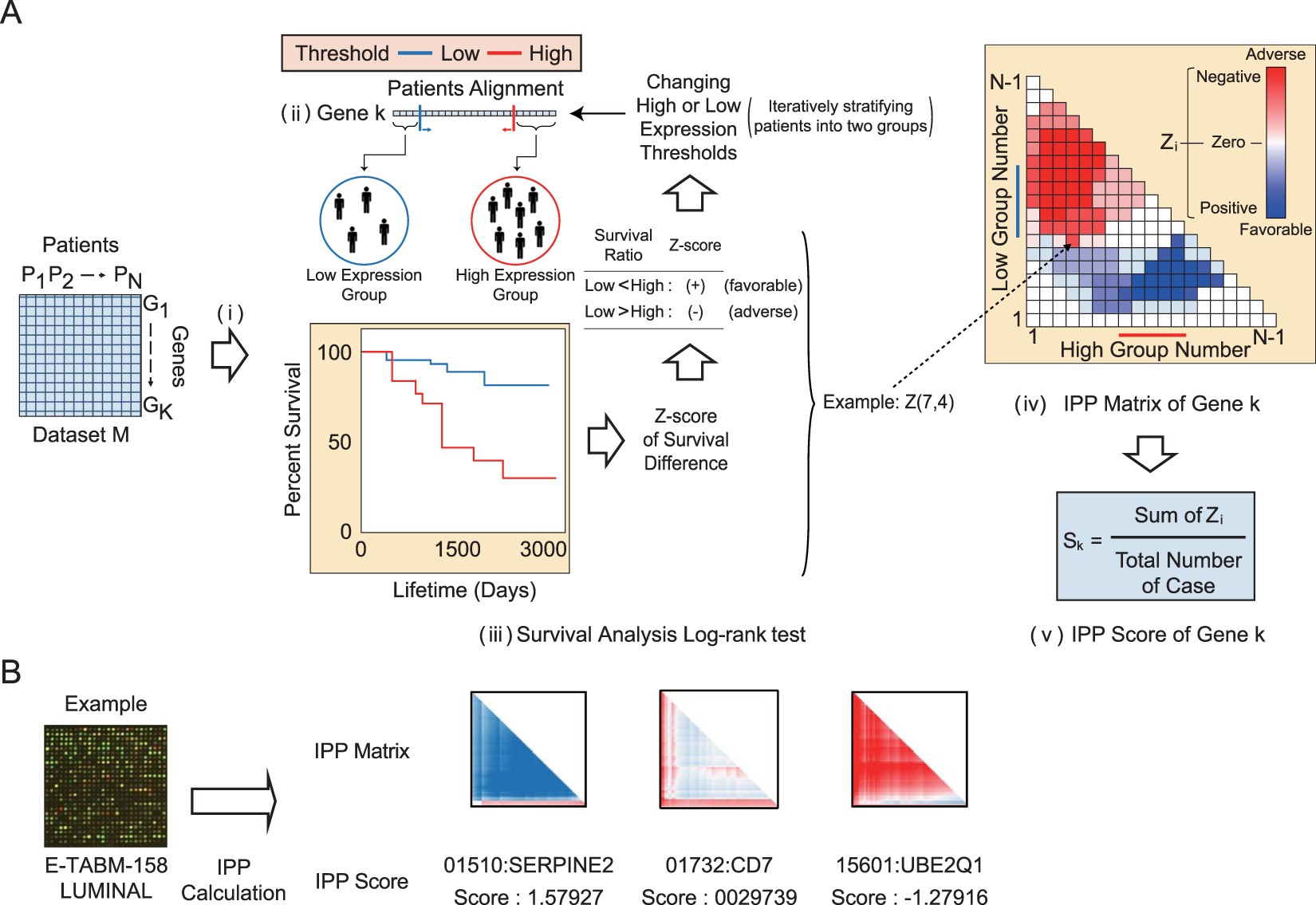
Robust method for identification of prognostic gene signatures from gene expression profiles | Scientific Reports

Distinct gene regulatory signatures of dominance rank and social bond strength in wild baboons | Philosophical Transactions of the Royal Society B: Biological Sciences
Identification of hub genes in prostate cancer using robust rank aggregation and weighted gene co-expression network analysis - Figure f3 | Aging
![MutRank: an R shiny web-application for exploratory targeted mutual rank-based coexpression analyses integrated with user-provided supporting information [PeerJ] MutRank: an R shiny web-application for exploratory targeted mutual rank-based coexpression analyses integrated with user-provided supporting information [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10264/1/fig-1-full.png)
MutRank: an R shiny web-application for exploratory targeted mutual rank-based coexpression analyses integrated with user-provided supporting information [PeerJ]

Dominance rank-associated gene expression is widespread, sex-specific, and a precursor to high social status in wild male baboons | PNAS
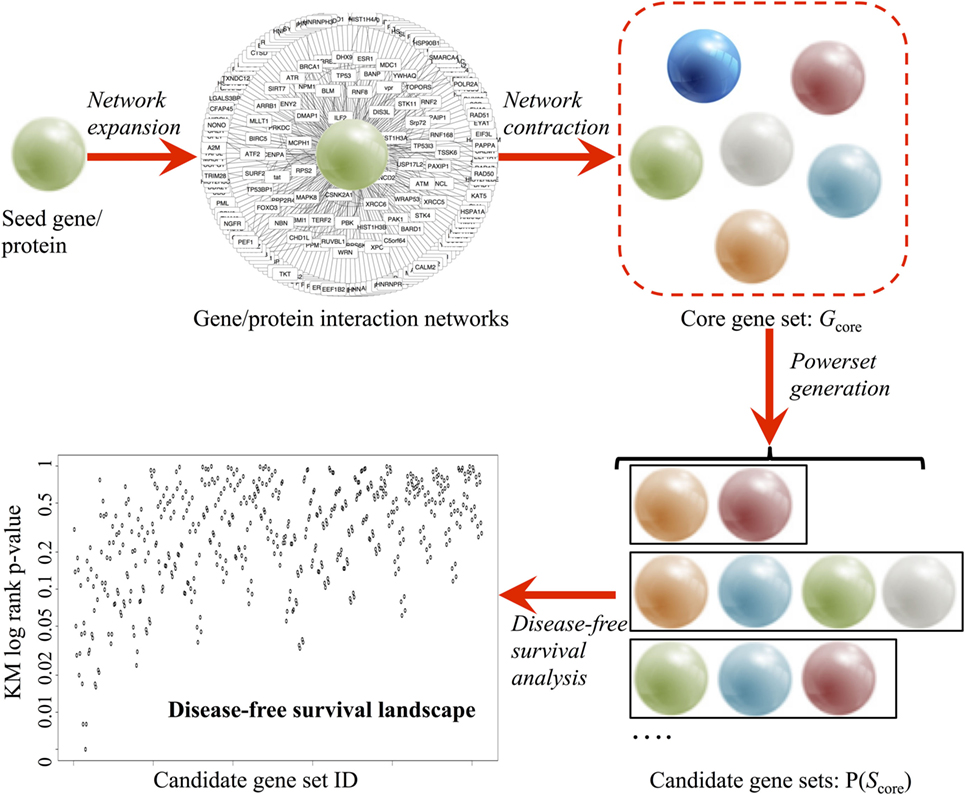
Frontiers | Combinatorial Ranking of Gene Sets to Predict Disease Relapse: The Retinoic Acid Pathway in Early Prostate Cancer